Orthomyxoviridae
| Orthomyxoviridae | |
|---|---|
| Virus classification | |
| Group: | Group V ((-)ssRNA) |
| Family: | Orthomyxoviridae |
| Genera | |
|
Influenzavirus A |
|
| Influenza (Flu) |
|---|
 |
| Types |
| Avian (A/H5N1 subtype) · Canine Equine · Swine (A/H1N1 subtype) |
| Vaccines |
| 2009 pandemic (Pandemrix) ACAM-FLU-A · Fluzone · Influvac Live attenuated (FluMist) · Optaflu |
| Treatment |
| Amantadine · Arbidol · Laninamivir Oseltamivir · Peramivir · Rimantadine Vitamin D · Zanamivir |
| Pandemics |
| 2009 · 1968–1969 Hong Kong · 1918 |
| Outbreaks |
| 2008 West Bengal 2007 Bernard Matthews H5N1 2007 Australian equine 2006 H5N1 India · 1976 swine flu |
| See also |
| Flu season · Influenza evolution Influenza research Influenza-like illness |
The Orthomyxoviridae (orthos, Greek for "straight"; myxa, Greek for "mucus")[1] are a family of RNA viruses that includes five genera: Influenzavirus A, Influenzavirus B, Influenzavirus C, Isavirus and Thogotovirus. A sixth has recently been described.[2] The first three genera contain viruses that cause influenza in vertebrates, including birds (see also avian influenza), humans, and other mammals. Isaviruses infect salmon; thogotoviruses infect vertebrates and invertebrates, such as mosquitoes and sea lice.[3][4][5]
The three genera of Influenzavirus, which are identified by antigenic differences in their nucleoprotein and matrix protein infect vertebrates as follows:
- Influenzavirus A cause of all flu pandemics and infect humans, other mammals and birds
- Influenzavirus B infect humans and seals
- Influenzavirus C infect humans and pigs
Contents |
Classification
In a phylogenetic-based taxonomy the "RNA viruses" includes the "negative-sense ssRNA viruses" which includes the Order "Mononegavirales", and the Family "Orthomyxoviridae" (among others). The genera-associated species and serotypes of Orthomyxoviridae are shown in the following table.
| Genus | Species (* indicates type species) | Serotypes or Subtypes | Hosts |
|---|---|---|---|
| Influenzavirus A | Influenza A virus* | H1N1, H1N2, H2N2, H3N1, H3N2, H3N8, H5N1, H5N2, H5N3, H5N8, H5N9, H7N1, H7N2, H7N3, H7N4, H7N7, H9N2, H10N7 | Human, pig, bird, horse |
| Influenzavirus B | Influenza B virus* | Human, seal | |
| Influenzavirus C | Influenza C virus* | Human, pig | |
| Isavirus | Infectious salmon anemia virus* | Atlantic salmon | |
| Thogotovirus | Thogoto virus* | Tick, mosquito, mammal (including human) | |
| Dhori virus | Batken virus, Dhori virus | ||
| Quaranfil virus, Johnston Atoll virus, Lake Chad virus |
Types
There are three genera of influenza virus: Influenzavirus A, Influenzavirus B and Influenzavirus C. Each genus includes only one species, or type: Influenza A virus, Influenza B virus, and Influenza C virus, respectively. Influenza A and C infect multiple species, while influenza B almost exclusively infects humans.[6][7]
Influenza A
Influenza A viruses are further classified, based on the viral surface proteins hemagglutinin (HA or H) and neuraminidase (NA or N)... Sixteen H subtypes (or serotypes) and nine N subtypes of influenza A virus have been identified.

Further variation exists; thus, specific influenza strain isolates are identified by a standard nomenclature specifying virus type, geographical location where first isolated, sequential number of isolation, year of isolation, and HA and NA subtype.[8][9]
Examples of the nomenclature are:
- A/Brisbane/59/2007 (H1N1)
- A/Moscow/10/99 (H3N2)
The type A viruses are the most virulent human pathogens among the three influenza types and causes the most severe disease. The serotypes that have been confirmed in humans, ordered by the number of known human pandemic deaths, are:
- H1N1 caused "Spanish Flu" in 1918, "Swine flu" in 2009[10]
- H2N2 caused "Asian Flu".
- H3N2 caused "Hong Kong Flu".
- H5N1 is a pandemic threat.
- H7N7 has unusual zoonotic potential.[11]
- H1N2 is endemic in humans and pigs.
- H9N2, H7N2, H7N3, H10N7.
| Name | Year | Deaths (millions) | Subtype involved |
|---|---|---|---|
| Asiatic (Russian) Flu | 1889-90 | 1 | possibly H2N2 |
| Spanish Flu | 1918-20 | 40 | H1N1 |
| Asian Flu | 1957-58 | 1-1.5 | H2N2 |
| Hong Kong Flu | 1968-69 | 0.75 | H3N2 |
| Swine Flu | 2009 - | ... | H1N1 |
Influenza B
Influenza B virus is almost exclusively a human pathogen, and is less common than influenza A. The only other animal known to be susceptible to influenza B infection is the seal.[13] This type of influenza mutates at a rate 2-3 times lower than type A[14] and consequently is less genetically diverse, with only one influenza B serotype.[6] As a result of this lack of antigenic diversity, a degree of immunity to influenza B is usually acquired at an early age. However, influenza B mutates enough that lasting immunity is not possible.[15] This reduced rate of antigenic change, combined with its limited host range (inhibiting cross species antigenic shift), ensures that pandemics of influenza B do not occur.[16]
Influenza C
The influenza C virus infects humans and pigs, and can cause severe illness and local epidemics.[17] However, influenza C is less common than the other types and usually seems to cause mild disease in children.[18][19]
Virology
Morphology
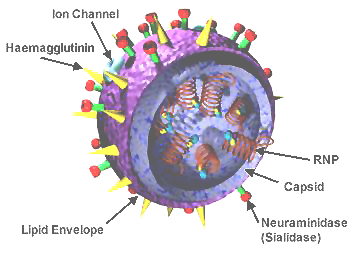
The virion is pleomorphic, the envelope can occur in spherical and filamentous forms. In general the virus's morphology is spherical with particles 50 to 120 nm in diameter, or filamentous virions 20 nm in diameter and 200 to 300 (-3000) nm long. There are some 500 distinct spike-like surface projections of the envelope each projecting 10 to 14 nm from the surface with some types (i.e. hemagglutinin esterase (HEF)) densely dispersed over the surface, and with others (i.e. hemagglutinin (HA)) spaced widely apart.
The major glycoprotein (HA) is interposed irregularly by clusters of neuraminidase (NA), with a ratio of HA to NA of about 4-5 to 1.
Lipoprotein membranes enclose the nucleocapsids; nucleoproteins of different size classes with a loop at each end; the arrangement within the virion is uncertain. The nucleocapsids are filamentous and fall in the range of 50 to 130 nm long and 9 to 15 nm in diameter. They have a helical symmetry.
Genome
Viruses of this family contain 7 to 8 segments of linear negative-sense single stranded RNA.
The total genome length is 12000-15000 nucleotides (nt). The largest segment 2300-2500 nt; of second largest 2300-2500 nt; of third 2200-2300 nt; of fourth 1700-1800 nt; of fifth 1500-1600 nt; of sixth 1400-1500 nt; of seventh 1000-1100 nt; of eighth 800-900 nt. Genome sequence has terminal repeated sequences; repeated at both ends. Terminal repeats at the 5'-end 12-13 nucleotides long. Nucleotide sequences of 3'-terminus identical; the same in genera of same family; most on RNA (segments), or on all RNA species. Terminal repeats at the 3'-end 9-11 nucleotides long. Encapsidated nucleic acid is solely genomic. Each virion may contain defective interfering copies.
Structure
The following applies for Influenza A viruses, although other influenza strains are very similar in structure:[20]
The influenza A virus particle or virion is 80-120 nm in diameter and usually roughly spherical, although filamentous forms can occur.[21] Unusually for a virus, the influenza A genome is not a single piece of nucleic acid; instead, it contains eight pieces of segmented negative-sense RNA (13.5 kilobases total), which encode 11 proteins (HA, NA, NP, M1, M2, NS1, NEP, PA, PB1, PB1-F2, PB2).[22] The best-characterised of these viral proteins are hemagglutinin and neuraminidase, two large glycoproteins found on the outside of the viral particles. Neuraminidase is an enzyme involved in the release of progeny virus from infected cells, by cleaving sugars that bind the mature viral particles. By contrast, hemagglutinin is a lectin that mediates binding of the virus to target cells and entry of the viral genome into the target cell.[23] The hemagglutinin (H) and neuraminidase (N) proteins are targets for antiviral drugs.[24] These proteins are also recognised by antibodies, i.e. they are antigens.[12] The responses of antibodies to these proteins are used to classify the different serotypes of influenza A viruses, hence the H and N in H5N1.
Life cycle

Typically, influenza is transmitted from infected mammals through the air by coughs or sneezes, creating aerosols containing the virus, and from infected birds through their droppings. Influenza can also be transmitted by saliva, nasal secretions, feces and blood. Infections occur through contact with these bodily fluids or with contaminated surfaces. Flu viruses can remain infectious for about one week at human body temperature, over 30 days at 0 °C (32 °F), and indefinitely at very low temperatures (such as lakes in northeast Siberia). They can be inactivated easily by disinfectants and detergents.[25][26][27]
The viruses bind to a cell through interactions between its hemagglutinin glycoprotein and sialic acid sugars on the surfaces of epithelial cells in the lung and throat (Stage 1 in infection figure).[28] The cell imports the virus by endocytosis. In the acidic endosome, part of the haemagglutinin protein fuses the viral envelope with the vacuole's membrane, releasing the viral RNA (vRNA) molecules, accessory proteins and RNA-dependent RNA polymerase into the cytoplasm (Stage 2).[29] These proteins and vRNA form a complex that is transported into the cell nucleus, where the RNA-dependent RNA transcriptase begins transcribing complementary positive-sense cRNA (Steps 3a and b).[30] The cRNA is either exported into the cytoplasm and translated (step 4), or remains in the nucleus. Newly-synthesised viral proteins are either secreted through the Golgi apparatus onto the cell surface (in the case of neuraminidase and hemagglutinin, step 5b) or transported back into the nucleus to bind vRNA and form new viral genome particles (step 5a). Other viral proteins have multiple actions in the host cell, including degrading cellular mRNA and using the released nucleotides for vRNA synthesis and also inhibiting translation of host-cell mRNAs.[31]
Negative-sense vRNAs that form the genomes of future viruses, RNA-dependent RNA transcriptase, and other viral proteins are assembled into a virion. Hemagglutinin and neuraminidase molecules cluster into a bulge in the cell membrane. The vRNA and viral core proteins leave the nucleus and enter this membrane protrusion (step 6). The mature virus buds off from the cell in a sphere of host phospholipid membrane, acquiring hemagglutinin and neuraminidase with this membrane coat (step 7).[32] As before, the viruses adhere to the cell through hemagglutinin; the mature viruses detach once their neuraminidase has cleaved sialic acid residues from the host cell.[28] After the release of new influenza virus, the host cell dies.
Since RNA proofreading enzymes are absent, the RNA-dependent RNA transcriptase makes a single nucleotide insertion error roughly every 10 thousand nucleotides, which is the approximate length of the influenza vRNA. Hence, nearly every newly-manufactured influenza virus will contain a mutation in its genome.[33] The separation of the genome into eight separate segments of vRNA allows mixing (reassortment) of the genes if more than one variety of influenza virus has infected the same cell (superinfection). The resulting alteration in the genome segments packaged in to viral progeny confers new behavior, sometimes the ability to infect new host species or to overcome protective immunity of host populations to its old genome (in which case it is called an antigenic shift).[12]
Viability and disinfection
Mammalian influenza virus tend to be labile, but can survive several hours in mucus.[34] Avian influenza virus can survive for 100 days in distilled water at room temperature, and 200 days at 17 °C (63 °F). The avian virus is inactivated more quickly in manure, but can survive for up to 2 weeks in feces on cages. Avian influenza viruses can survive indefinitely when frozen.[34] Influenza viruses are susceptible to bleach, 70% ethanol, aldehydes, oxidizing agents, and quaternary ammonium compounds. They are inactivated by heat of 133 °F (56 °C) for minimum of 60 minutes, as well as by low pH <2.[34]
References
- ↑ International Committee on Taxonomy of Viruses Index of Viruses - Orthomyxoviridae (2006). In: ICTVdB - The Universal Virus Database, version 4. Büchen-Osmond, C (Ed), Columbia University, New York, USA.
- ↑ Presti RM, Zhao G, Beatty WL, Mihindukulasuriya KA, Travassos da Rosa AP, Popov VL, Tesh RB, Virgin HW, Wang D(2009) Quaranfil, Johnston Atoll and Lake Chad viruses are novel members of the family Orthomyxoviridae. J Virol.
- ↑ Jones LD, Nuttall PA (1989). "Non-viraemic transmission of Thogoto virus: influence of time and distance". Trans. R. Soc. Trop. Med. Hyg. 83 (5): 712–4. doi:10.1016/0035-9203(89)90405-7. PMID 2617637.
- ↑ Barry Ely (1999). "Infectious Salmon Anaemia". Mill Hill Essays. National Institute for Medical Research. http://www.nimr.mrc.ac.uk/MillHillEssays/1999/isa.htm. Retrieved 2007-09-14.
- ↑ Raynard RS, Murray AG, Gregory A (2001). "Infectious salmon anaemia virus in wild fish from Scotland". Dis. Aquat. Org. 46 (2): 93–100. doi:10.3354/dao046093. PMID 11678233.
- ↑ 6.0 6.1 Hay A, Gregory V, Douglas A, Lin Y (Dec 29 2001). "The evolution of human influenza viruses" (PDF). Philos Trans R Soc Lond B Biol Sci 356 (1416): 1861–70. doi:10.1098/rstb.2001.0999. PMID 11779385. PMC 1088562. http://www.journals.royalsoc.ac.uk/media/hf0bujxwvrcxd7nwdrwq/contributions/l/x/y/v/lxyv2p8w45geev90.pdf.
- ↑ "Avian Influenza (Bird Flu)". Centers for Disease Control and Prevention. http://www.cdc.gov/flu/avian/. Retrieved 2007-09-15.
- ↑ Atkinson W, Hamborsky J, McIntyre L, Wolfe S, ed (2007). Epidemiology and Prevention of Vaccine-Preventable Diseases (10th ed.). Washington DC: Centers for Disease Control and Prevention. http://www.cdc.gov/vaccines/pubs/pinkbook/pink-chapters.htm.
- ↑ "Avian Influenza (Bird Flu): Implications for Human Disease". Center for Infectious Disease Research & Policy, University of Minnesota. 2007-06-27. http://www.cidrap.umn.edu/cidrap/content/influenza/avianflu/biofacts/avflu_human.html. Retrieved 2007-09-14.
- ↑ Wang TT, Palese P. Cell. 2009 Jun 12;137(6):983-5. PMID: 19524497
- ↑ Fouchier R, Schneeberger P, Rozendaal F, Broekman J, Kemink S, Munster V, Kuiken T, Rimmelzwaan G, Schutten M, Van Doornum G, Koch G, Bosman A, Koopmans M, Osterhaus A (2004). "Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome". Proc Natl Acad Sci USA 101 (5): 1356–61. doi:10.1073/pnas.0308352100. PMID 14745020.
- ↑ 12.0 12.1 12.2 Hilleman M (Aug 19 2002). "Realities and enigmas of human viral influenza: pathogenesis, epidemiology and control". Vaccine 20 (25-26): 3068–87. doi:10.1016/S0264-410X(02)00254-2. PMID 12163258.
- ↑ Osterhaus A, Rimmelzwaan G, Martina B, Bestebroer T, Fouchier R (2000). "Influenza B virus in seals". Science 288 (5468): 1051–3. doi:10.1126/science.288.5468.1051. PMID 10807575.
- ↑ Nobusawa E, Sato K (April 2006). "Comparison of the mutation rates of human influenza A and B viruses". J Virol 80 (7): 3675–8. doi:10.1128/JVI.80.7.3675-3678.2006. PMID 16537638.
- ↑ Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y (March 1992). "Evolution and ecology of influenza A viruses". Microbiol. Rev. 56 (1): 152–79. PMID 1579108. PMC 372859. http://mmbr.asm.org/cgi/pmidlookup?view=long&pmid=1579108.
- ↑ Zambon M (November 1999). "Epidemiology and pathogenesis of influenza". J Antimicrob Chemother 44 (Suppl B): 3–9. doi:10.1093/jac/44.suppl_2.3. PMID 10877456. http://jac.oxfordjournals.org/cgi/reprint/44/suppl_2/3.
- ↑ Matsuzaki Y, Sugawara K, Mizuta K, Tsuchiya E, Muraki Y, Hongo S, Suzuki H, Nakamura K (2002). "Antigenic and genetic characterization of influenza C viruses which caused two outbreaks in Yamagata City, Japan, in 1996 and 1998". J Clin Microbiol 40 (2): 422–9. doi:10.1128/JCM.40.2.422-429.2002. PMID 11825952. PMC 153379. http://jcm.asm.org/cgi/pmidlookup?view=long&pmid=11825952.
- ↑ Matsuzaki Y, Katsushima N, Nagai Y, Shoji M, Itagaki T, Sakamoto M, Kitaoka S, Mizuta K, Nishimura H (May 1 2006). "Clinical features of influenza C virus infection in children". J Infect Dis 193 (9): 1229–35. doi:10.1086/502973. PMID 16586359.
- ↑ Katagiri S, Ohizumi A, Homma M (July 1983). "An outbreak of type C influenza in a children's home". J Infect Dis 148 (1): 51–6. PMID 6309999.
- ↑ International Committee on Taxonomy of Viruses descriptions of: Orthomyxoviridae Influenzavirus B Influenzavirus C
- ↑ International Committee on Taxonomy of Viruses. "The Universal Virus Database, version 4: Influenza A". http://www.ncbi.nlm.nih.gov/ICTVdb/ICTVdB/00.046.0.01.htm.
- ↑ Ghedin E, Sengamalay N, Shumway M, Zaborsky J, Feldblyum T, Subbu V, Spiro D, Sitz J, Koo H, Bolotov P, Dernovoy D, Tatusova T, Bao Y, St George K, Taylor J, Lipman D, Fraser C, Taubenberger J, Salzberg S (Oct 20 2005). "Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution". Nature 437 (7062): 1162–6. doi:10.1038/nature04239. PMID 16208317.
- ↑ Suzuki Y (2005). "Sialobiology of influenza: molecular mechanism of host range variation of influenza viruses". Biol Pharm Bull 28 (3): 399–408. doi:10.1248/bpb.28.399. PMID 15744059. http://www.jstage.jst.go.jp/article/bpb/28/3/399/_pdf.
- ↑ Wilson J, von Itzstein M (July 2003). "Recent strategies in the search for new anti-influenza therapies". Curr Drug Targets 4 (5): 389–408. doi:10.2174/1389450033491019. PMID 12816348.
- ↑ Suarez, D; Spackman E, Senne D, Bulaga L, Welsch A, Froberg K (2003). "The effect of various disinfectants on detection of avian influenza virus by real time RT-PCR". Avian Dis 47 (3 Suppl): 1091–5. doi:10.1637/0005-2086-47.s3.1091. PMID 14575118.
- ↑ Avian Influenza (Bird Flu): Implications for Human Disease. Physical characteristics of influenza A viruses. UMN CIDRAP.
- ↑ Flu viruses 'can live for decades' on ice, NZ Herald, November 30, 2006.
- ↑ 28.0 28.1 Wagner R, Matrosovich M, Klenk H (May-Jun 2002). "Functional balance between haemagglutinin and neuraminidase in influenza virus infections". Rev Med Virol 12 (3): 159–66. doi:10.1002/rmv.352. PMID 11987141.
- ↑ Lakadamyali M, Rust M, Babcock H, Zhuang X (Aug 5 2003). "Visualizing infection of individual influenza viruses". Proc Natl Acad Sci USA 100 (16): 9280–5. doi:10.1073/pnas.0832269100. PMID 12883000.
- ↑ Cros J, Palese P (September 2003). "Trafficking of viral genomic RNA into and out of the nucleus: influenza, Thogoto and Borna disease viruses". Virus Res 95 (1-2): 3–12. doi:10.1016/S0168-1702(03)00159-X. PMID 12921991.
- ↑ Kash J, Goodman A, Korth M, Katze M (July 2006). "Hijacking of the host-cell response and translational control during influenza virus infection". Virus Res 119 (1): 111–20. doi:10.1016/j.virusres.2005.10.013. PMID 16630668.
- ↑ Nayak D, Hui E, Barman S (December 2004). "Assembly and budding of influenza virus". Virus Res 106 (2): 147–65. doi:10.1016/j.virusres.2004.08.012. PMID 15567494.
- ↑ Drake J (May 1 1993). "Rates of spontaneous mutation among RNA viruses". Proc Natl Acad Sci USA 90 (9): 4171–5. doi:10.1073/pnas.90.9.4171. PMID 8387212.
- ↑ 34.0 34.1 34.2 http://www.cfsph.iastate.edu/Factsheets/pdfs/influenza.pdf, p. 7
35. ISBN 9783211808924 The Influenza Viruses Hoyle L 1968 Springer Verlag
External links
- Health-EU Portal EU work to prepare a global response to influenza.
- Influenza Research Database Database of influenza genomic sequences and related information.
- Swine Flu News (World View) Swine Flu News (World View)
- European Commission - Public Health EU coordination on Pandemic (H1N1) 2009
|
|||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||